Golden Gate Express II, Grand Island, Nebraska. 42 likes 338 were here. Golden Gate Assembly The efficient and seamless assembly of DNA fragments, commonly referred to as Golden Gate assembly, has its origins in 1996, when for the first time it was shown that multiple inserts could be assembled into a vector backbone using only the sequential or simultaneous activities of a single type IIS restriction enzyme and T4 DNA ligase.
IPSWICH, Mass., June 11, 2018 /PRNewswire/ - Some of today's most exciting molecular biology engineering — including the creation of customized CRISPR/Cas9 and TALEN constructs for genome editing — are led by synthetic biologists. These scientists are using methods such as Golden Gate Assembly. This molecular cloning technique uses a combination of Type IIS restriction enzymes and DNA ligase to assemble multiple DNA fragments simultaneously in a rapid 'one-pot' reaction.However, there are practical limitations to Golden Gate Assembly, which is highly dependent upon the efficient ligation of flanking four base pair overhangs with T4 DNA Ligase, where a difference in even one base may create unwanted ligation products.
As a result, current best practices for the number of DNA fragments that can be joined in a single reaction have been typically limited to ten fragments or less.A new ligase fidelity profile, developed and tested by researchers at New England Biolabs (NEB ®), used single molecule real-time sequencing to identify and quantify the proportions of mismatched-prone junction pairs during ligation. The assay accurately predicted junction fidelity for all 256 possible 4-base overhangs, allowing users to choose optimal Golden Gate ligation junctions when designing assemblies.' Without a proper understanding of ligase fidelity and bias, mismatched overhangs may be ligated — leading to the inclusion of mismatches, deletions or insertions into the final DNA vector assemblies,' said Gregory Lohman, Staff Scientist at NEB and author on the study. 'The application of our ligation fidelity data effectively informed our choice of junctions during Golden Gate Assembly, enabling a highly flexible design and successful assembly of more than 20 fragments in a single reaction. Therefore, we're making our ligase fidelity data publicly available to help scientists fine-tune their Golden Gate protocol.'
Additionally, NEB has introduced two new restriction enzymes — BsaI-HF ®v2 and Esp3I; BsaI-HFv2 has been re-engineered and optimized for Golden Gate Assembly and Esp3I is an isoschizomer of BsmBI.' NEB has an extensive history innovating in molecular biology enzymes and related technologies,' said Rebecca Kucera, Applications and Product Development Scientist at NEB. 'In addition to the ligase fidelity profile data, the availability of BsaI-HFv2, a key type IIS restriction enzyme, has resulted in improved performance for Golden Gate Assembly, with highly efficient reactions yielding 99% accuracy for 12-fragment assemblies and greater than 90% accuracy for 24-fragment assemblies.'
The ligase fidelity research is being conducted as part of a broader effort to explore the fidelity of other commercially available enzymes, including transcriptases, polymerases, and exo/endonucleases, further enhancing standard molecular biology techniques.' NEB's research and development efforts will continue to focus on providing industry-leading reaction components for Golden Gate assembly, with the long-term vision of enabling efficient assembly of 50 fragments in a single reaction,' said Lohman.For more information, visit.About New England BiolabsEstablished in the mid 1970's, New England Biolabs, Inc. (NEB) is the industry leader in the discovery and production of enzymes for molecular biology applications and now offers the largest selection of recombinant and native enzymes for genomic research. NEB continues to expand its product offerings into areas related to PCR, gene expression, sample preparation for next generation sequencing, synthetic biology, glycobiology, epigenetics and RNA analysis. Additionally, NEB is focused on strengthening alliances that enable new technologies to reach key market sectors, including molecular diagnostics development. New England Biolabs is a privately held company, headquartered in Ipswich, MA, and has extensive worldwide distribution through a network of exclusive distributors, agents and seven subsidiaries located in Canada, China, France, Germany, Japan, Singapore and the UK. For more information about New England Biolabs visitNEB ® and NEW ENGLAND BIOLABS ® are registered trademarks of New England Biolabs, Inc.SOURCE New England Biolabs Related Links.
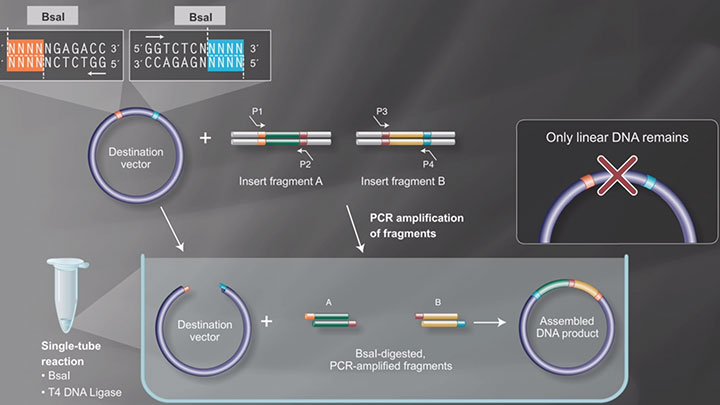
Continuing toward our goal of making cloning painless, we’re releasing a new method in our assembly wizard that many scientists have requested: Golden Gate Assembly.We built this tool in conjunction with scientists from, who offered their scientific expertise to help us formulate how the wizard should work. Stay tuned for further updates about how our partnership with NEB will create better tools for researchers.Golden Gate uses Type IIs restriction enzymes which make 4 bp sticky ends next to their recognition sites. As long as the sticky ends can’t improperly anneal, multiple fragments can combined in a single pot with high efficiency.Selecting appropriate sticky ends by hand is a time-consuming and error-prone process. Our assembly wizard automates sticky end design and supports more complicated techniques such as including spacers between fragments and site-directed mutagenesis.Below is a quick description of how to use the tool. For a complete description, check out our.
As a reminder, our assembly wizard also supports cloning through Gibson and digest-and-ligate methods. Creating a new assemblyTo start using the assembly wizard, open any sequence in and create a new assembly from the “Assembly Wizard” button at the bottom right.You can set each fragment from a pair of type IIs cut sites or from any sequence selection. In the latter case, the wizard automatically generates primers that introduce recognition sites and valid sticky ends for the selected enzymes. Add spacer sequences using the icon next to each fragment.When you’re done selecting fragments, click the Assemble button to generate the output plasmid. Viewing and adjusting the output plasmidThe Assembly tab in the output plasmid contains information about the primers that were designed.
Buttons at the top help you export the primers and print a summary for your lab notebook.To add point mutations, simply select any bases covered by a primer on the sequence map and type to replace. The relevant primers are automatically updated to incorporate the change, and mismatches with the parent sequence are labeled.
Benchling versions the sequence and primers so you can undo any edit you make.When you’re done adjusting the design, select a folder where the primers should be saved and click “Finalize” from the Assembly tab. This will fully save your plasmid and enable all of Benchling’s features on it, such as sequence alignment for verification.As always, feedback on this new feature is welcome. If you have suggestions or would like to see a new assembly technique incorporated, drop us a line at. Stay tuned for further updates, such as combinatorial assembly that allows you to set a library of parts for each fragment and generate all possible combinations in a single click.Don’t have an account? Sign up for free at.
